Functional Brain Imaging combining DOI and fMRI
1. About
Diffuse Optical Imaging
Diffuse Optical Imaging (DOI)
is a relatively new method used to image blood volume and oxygen saturation in
vivo. It uses near infrared light and
has the advantages of being low cost and portable. The properties of near infrared light in
biological tissue make diffuse optical imaging techniques very successful. The absorption coefficient (µa)
depends on the total hemoglobin concentration and oxygenation level within the
tissue; therefore, calculating µa provides useful information about
the physiological conditions of the tissue.
For instance, during the last few years DOI has been tested for
application to breast cancer imaging, brain oxygenation, brain trauma, and
brain function.
2. Motivation
Various methods are known for
producing functional brain images.
Functional Magnetic Resonance Imaging (fMRI) provides good spatial
resolution but poor temporal resolution whereas DOI exhibits excellent temporal
resolution but is penalized by the highly scattering superficial head layers
that make it difficult for light to reach the brain. Therefore, the DOI signal scattered back from
the brain and detected by the optodes placed on the scalp is fairly weak.
Intuitively, a multi-modality
method should provide more accurate functional images of the brain because it
combines the strengths of diverse methods.
The current project aims to thoroughly explore the potential of a
dual-modality system that uses MRI to spatially guide DOI.
3.
Dual-modality method and preliminary results
The idea is to use fMRI to
localize the region of the cortex where there is activity during the
performance of a task (e.g. finger tapping, visual stimuli, etc.) and to
linearly combine the data acquired using fMRI with that gathered simultaneously
by DOI.
Using a soft spatial prior
provided by fMRI we add information about the position of the activation region
in the brain. By weighting such prior we
can make it more or less relevant in the reconstruction process. For example, if we believe that DOI can
localize the activation region by itself, we can assign a zero-weight to the
fMRI prior. On the other hand, if the
activation region is in deeper layers and therefore hardly detectable by our
optical system, we can increase the contribution of the spatial prior in the
reconstruction process.
Unfortunately, the soft prior
added to the Tikhonov functional is not strong enough to significantly improve
the restored contrast image. Since the problem is extremely underdetermined
(84,864 unknowns and 114 measurements), the restoration algorithm is more
likely to find activation in the regions where the detectors are most sensitive
to absorption changes. Therefore, activation is typically found at the
superficial layers (note that light intensity decays exponentially with depth).
Figure 1 shows the head model
and the locations of maximal sensitivity to absorption in the cortex (white
crosses) of each optode (numbered in black) for a given coronal slice.
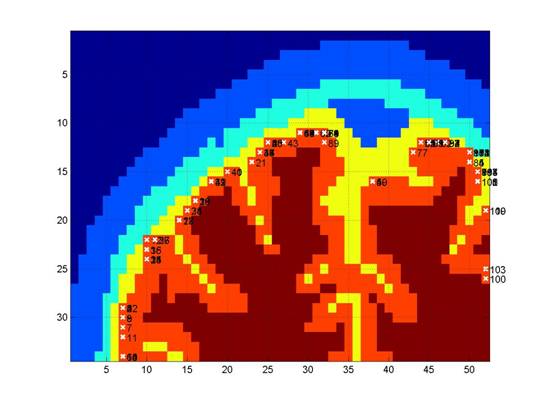
Figure 1
In order to improve
localization of the activated region it is necessary to reduce the number of
voxels involved in the restoration process. A simple way to acquire information
on the inactive region (such as scalp and skull tissue types, where brain
activation will not induce absorption coefficient changes) is to perform an MR
anatomical scan and assign zero value to the voxels corresponding to scalp and
skull in the imaging matrix calculated from DOI. We call the introduction of
such a hard constraint to the DOI forward model the scalp-skull prior (or Hard Brain in figure 2a and figure 2b).
Figure 2 and figure 3
demonstrate the improvement of the dual-modality method over each technique
alone: figure 2a and figure 2b show coronal slices of the brain reconstructed
using the Tikhonov inverse method and a hard prior (the second column shows the
restoration using the scalp-skull prior, whereas the third column shows the
result calculated by adding the cortical prior as in [1]). Figure 2a presents an example of two coronal
slices where the use of the scalp-skull prior produces better images than using
the cortical prior; figure 2b, on the other hand, shows a case where the use of
the cortical prior generates more accurate reconstructions. The columns and rows in figure 2 correspond
to different restoration modality and coronal slices, respectively. The first
column shows the true simulated activation region, the second column shows the
restoration obtained using the scalp-skull prior, and the third column shows
the results calculated using the cortical prior.
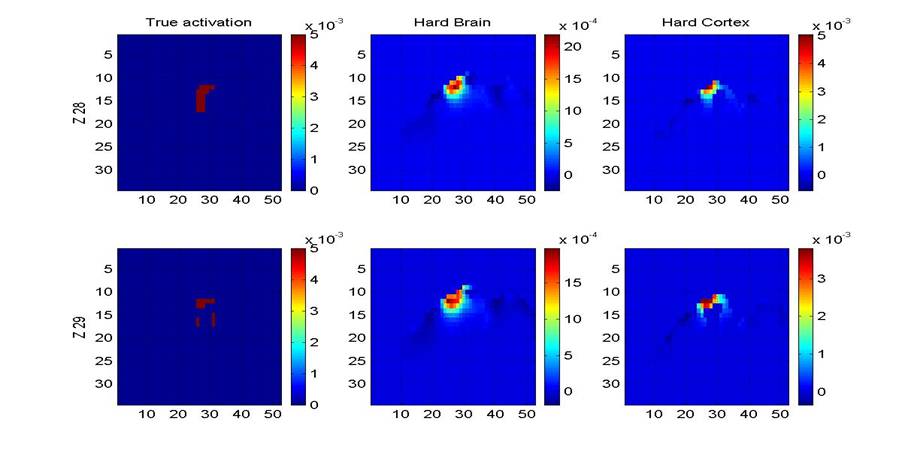
Figure 2a
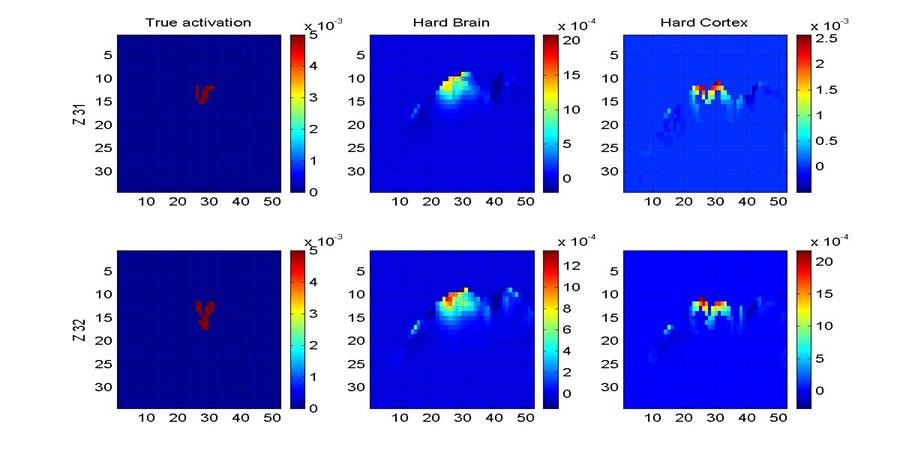
Figure 2b
Figure 3 compares a simple
Tikhonov reconstruction with a Tikhonov restoration using an fMRI hard prior
(i.e. forcing activation to be found in the location identified by the fMRI
data). The center column of the figure shows how powerful such a prior is and
how much information is therefore lost. Ideally we would like to find a compromise
that uses as much information as possible and preserves it through the
reconstruction process but at the same time finds the activation region
position with the greatest possible accuracy.
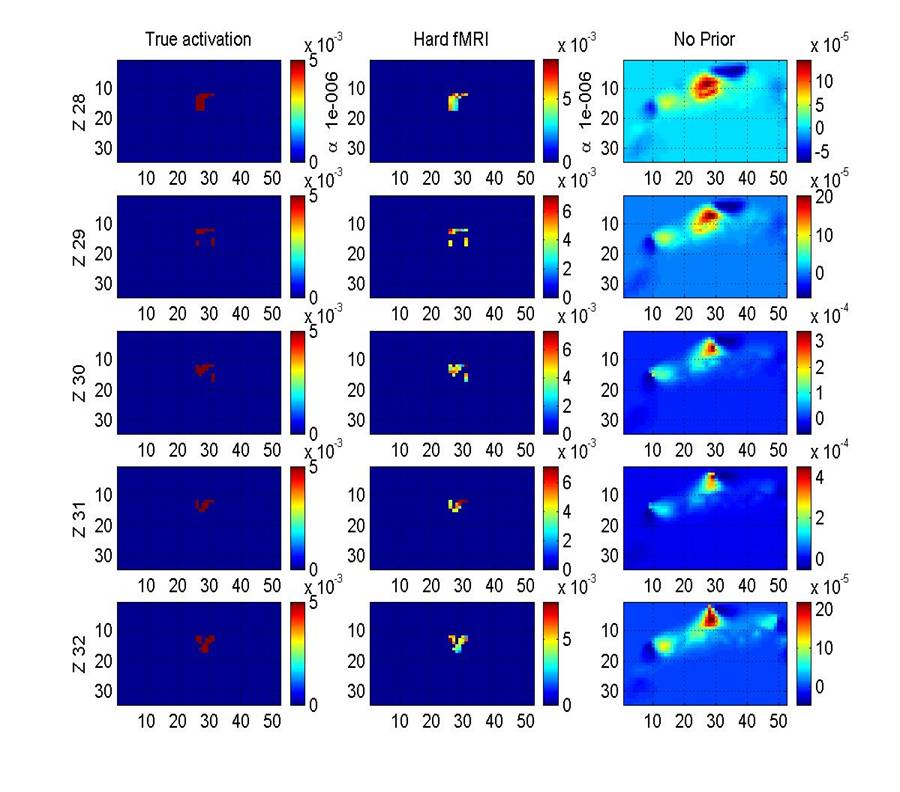
Figure 3
4. Summary
The above data showed that
using a spatial prior clearly improves the localization of the activation
region in the brain. The tests were
performed using only one temporal frame.
Combining all the temporal frames will express more accurately the
temporal evolution of the activation due to task performance and the
localization of the activation region in the brain. The localization, in particular, was improved
by the contribution of the MRI data.
The use of a scalp-skull
prior (i.e. adding information on locations of non active regions) instead of a
hard cortical prior (i.e. forcing sensitivity to µa changes to be
only in the cortex) has two main advantages: it decreases the prior information
used in the restoration process and it reduces the computation time required to
calculate the prior. This is because
computing the hard prior necessitates acquiring a full anatomical MR image of
the subject head and segmenting it into the various tissue types, whereas
computing the scalp-skull prior involves the performance of a preliminary MR
anatomical scan without segmentation that is extremely less computationally
expensive.
5. References
[1]
D. Boas and A. Dale, “A simulation study of MRI guided cortically constrained
diffuse optical tomography of human brain function” (2004)
Researchers
Anna Custo custo[at]csail.mit.edu
David Boas dboas[at]nmr.mgh.harvard.edu
Eric Grimson welg[at]csail.mit.edu
William Wells
sw[at]csail.mit.edu
John Fisher fisher[at]csail.mit.edu
Back
to the Medical Vision Group page.
Back
to the MIT CSAIL page.
Last updated
custo[at]csail.mit.edu